The Beer-Lambert law relates the concentration, $c$, of a substance in a solution sample to the intensity of light transmitted through the sample, $I_\mathrm{t}$ across a given path length, $l$, at a given wavelength, $\lambda$: $$ I_\mathrm{t} = I_0 e^{-\alpha c l}, $$ where $I_0$ is the incident light intensity and $\alpha$ is the absorption coefficient at $\lambda$.
Given a series of measurements of the fraction of light transmitted, $I_\mathrm{t}/I_0$, $\alpha$ may be determined through a least-squares fit to the straight line: $$ y = \ln\frac{I_\mathrm{t}}{I_0} = -\alpha c l. $$ Although this line passes through the origin ($y=0$ for $c=0$), we will fit the more general linear relationship: $$ y = mc + k $$ where $m = -\alpha l$, and verify that $k$ is close to zero.
Given a sample with path length $l = 0.8\;\mathrm{cm}$, the following data were measured for $I_\mathrm{t}/I_0$ at five different concentrations:
| $c\;/\mathrm{M}$ | $I_\mathrm{t}/I_0$ |
|---|---|
| 0.4 | 0.886 |
| 0.6 | 0.833 |
| 0.8 | 0.784 |
| 1.0 | 0.738 |
| 1.2 | 0.694 |
The matrix form of the least squares equation to be solved is
\begin{align*}
\left(\begin{array}{ll}
c_1 & 1\\
c_2 & 1\\
c_3 & 1\\
c_4 & 1\\
c_5 & 1\\
\end{array}\right)
\left(\begin{array}{l}
m \\ k
\end{array}\right)
=
\left(\begin{array}{ll}
T_1\\
T_2\\
T_3\\
T_4\\
T_5\\
\end{array}\right)
\end{align*}
where $T = \ln(I_\mathrm{t}/I_0)$. The code below determines $m$ and hence $\alpha$ using np.linalg.lstsq:
import numpy as np
import matplotlib.pyplot as plt
# Path length, cm
path = 0.8
# The data: concentrations (M) and It/I0
c = np.array([0.4, 0.6, 0.8, 1.0, 1.2])
It_over_I0 = np.array([ 0.891 , 0.841, 0.783, 0.744, 0.692])
n = len(c)
A = np.vstack((c, np.ones(n))).T
T = np.log(It_over_I0)
x, resid, _, _ = np.linalg.lstsq(A, T, rcond=None)
m, k = x
alpha = - m / path
print('alpha = {:.3f} M-1.cm-1'.format(alpha))
print('k', k)
print('rms residual = ', np.sqrt(resid[0]))
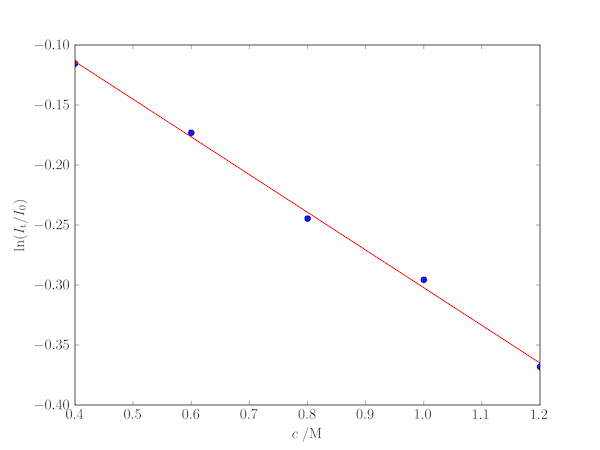
plt.plot(c, T, 'o')
plt.plot(c, m*c + k)
plt.xlabel('$c\;/\mathrm{M}$')
plt.ylabel('$\ln(I_\mathrm{t}/I_0)$')
plt.show()
Here, _ is the dummy variable name conventionally given to an object we do not need to store or use.
The output produces a best fit value of $\alpha=0.393\;\mathrm{M^{-1}\,cm^{-1}}$ and a value of $k$ compatible with experimental error:
alpha = 0.393 M-1.cm-1
k 0.0118109033334
rms residual = 0.0096843591966